Abstract
Autoregressive models for molecular graph generation typically operate on flattened sequences of atoms and bonds, discarding the rich multi-scale structure inherent to molecules. We introduce MOSAIC (Multi-scale Organization via Structural Abstraction In Composition), a framework that lifts autoregressive generation from flat token walks to compositional, hierarchy-aware sequences. MOSAIC provides a unified three-stage pipeline: (1) hierarchical coarsening that recursively groups atoms into motif-like clusters using graph-theoretic methods (spectral clustering, hierarchical agglomerative clustering, and motif-aware variants), (2) structured tokenization that serializes the resulting multi-level hierarchy into sequences that explicitly encode parent-child relationships, partition boundaries, and edge connectivity at every level, and (3) autoregressive generation with a standard Transformer decoder that learns to produce these structured sequences. We evaluate MOSAIC on the MOSES and COCONUT molecular benchmarks, comparing four tokenization schemes of increasing hierarchical expressiveness. Our experiments show that hierarchy-aware tokenizations improve chemical validity and structural diversity over flat baselines while enabling control over generated substructures. MOSAIC provides a principled, modular foundation for structure-aware molecular generation.
Introduction
Molecular graphs are a natural representation for drug discovery and property prediction: nodes are atoms, edges are bonds, and validity and function depend heavily on recurring substructures — rings, functional groups, and their connectivity. While discriminative graph neural networks excel at predicting properties from such graphs, generating novel, valid molecules remains challenging. Autoregressive transformers, which have scaled successfully in language and code, offer an attractive backbone for graph generation when graphs are first tokenized into sequences. Existing tokenization schemes typically flatten the graph into a single linear order — a random walk or depth-first traversal — so that a causal transformer can predict the next token. These flat walks capture local adjacency and some long-range edges via back-references, but they do not expose the graph’s compositional hierarchy: which atoms belong to which ring or functional group, and how those units connect. The model must therefore infer motif structure implicitly from the token stream, which can lead to broken rings, inconsistent motif statistics, and limited control over high-level chemistry.
MOSAIC addresses this gap by introducing an explicit hierarchical graph (H-graph) abstraction and a three-stage pipeline for autoregressive molecular generation. First, we coarsen the molecular graph into a set of communities and inter-community edges, optionally using domain knowledge (ring and functional-group detection) to guarantee motif cohesion. Second, we tokenize the H-graph into a flat sequence using a coarse-before-fine ordering: the model sees community-level structure before atom-level detail, mirroring the compositional nature of chemistry. Third, we flatten the generated token sequence back into a molecular graph. All tokenization schemes support a lossless roundtrip from graph to tokens and back.
Pipeline Overview
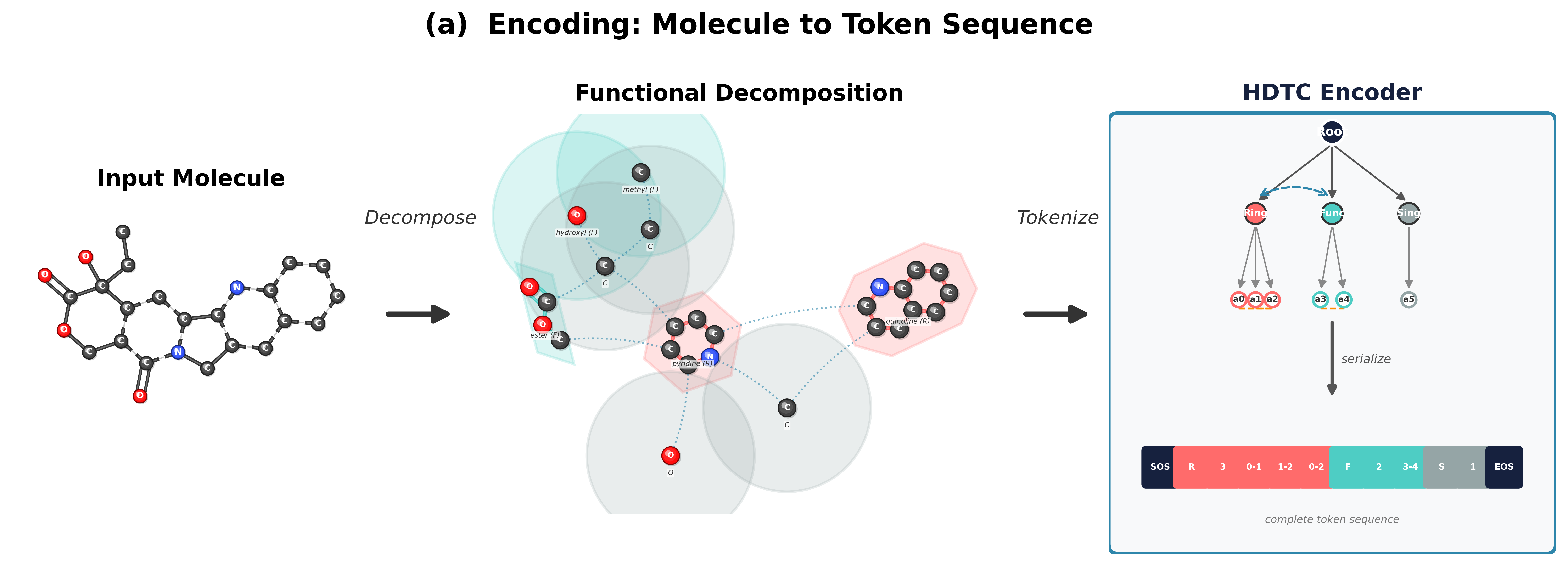
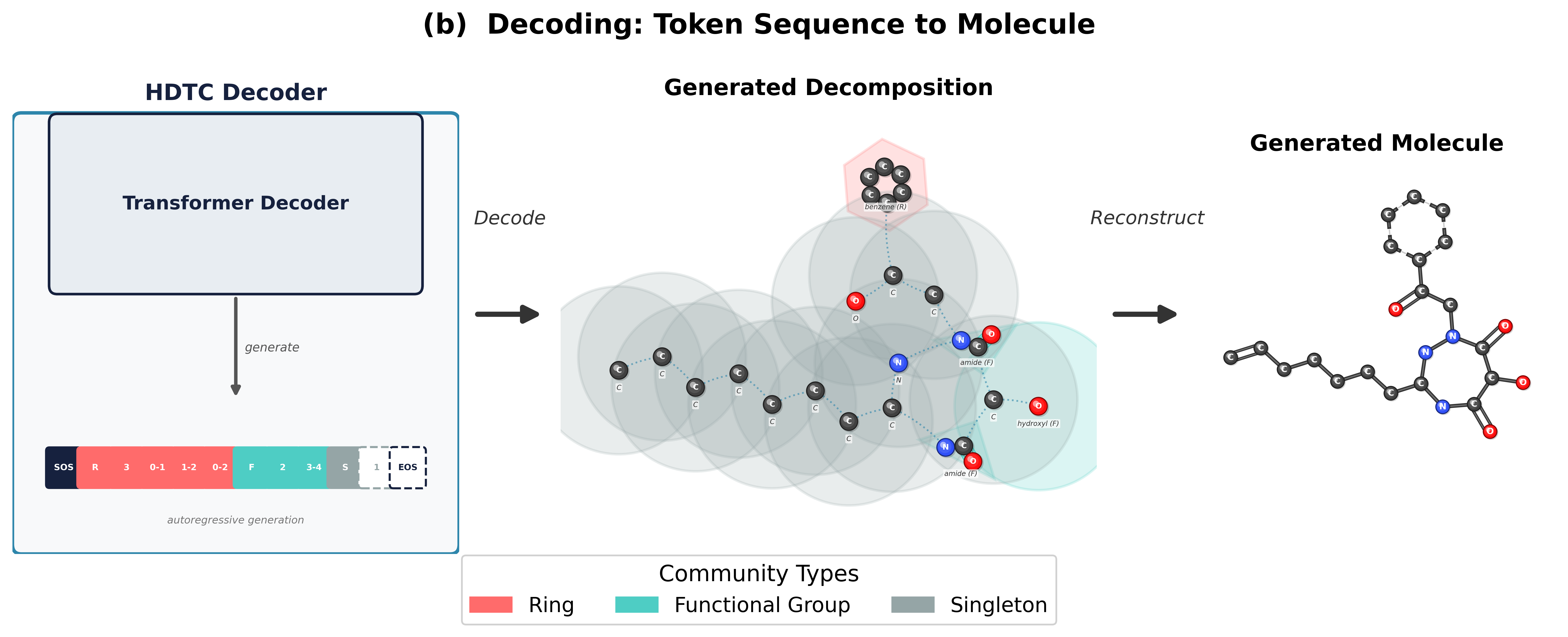
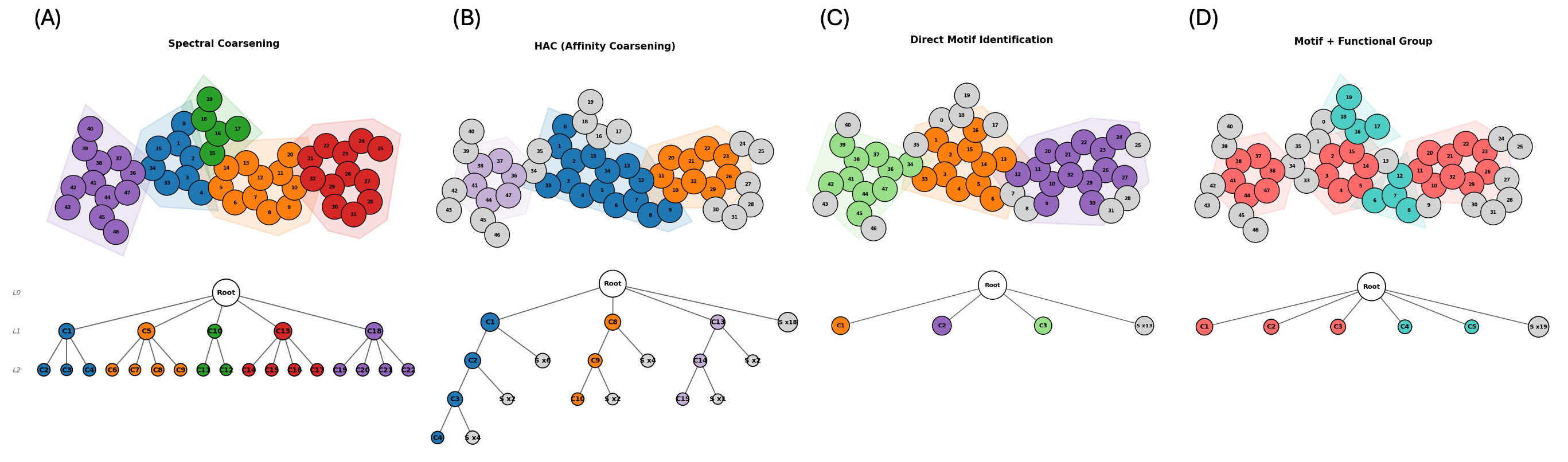
Coarsening Strategies
The coarsening step determines how atoms are grouped into communities. We explore two families: flexible methods that optimize a graph-theoretic objective without domain knowledge, and constraint methods that leverage chemical structure to define communities. Spectral clustering uses the eigenvectors of the graph Laplacian to identify natural clusters, selecting the number of partitions that maximizes Newman–Girvan modularity. Hierarchical agglomerative clustering (HAC) takes a bottom-up approach, iteratively merging the most similar adjacent nodes based on bond-type-weighted distances, which tends to preserve local chemical neighborhoods. Motif-aware coarsening (MC) first identifies known chemical motifs (rings, functional groups) via SMARTS pattern matching, then clusters the remaining atoms with spectral or HAC methods. This ensures that chemically meaningful substructures are preserved as intact units. MC with functional groups (MC+FG) extends this with an expanded library of functional-group patterns for finer-grained control over which substructures remain intact during coarsening.
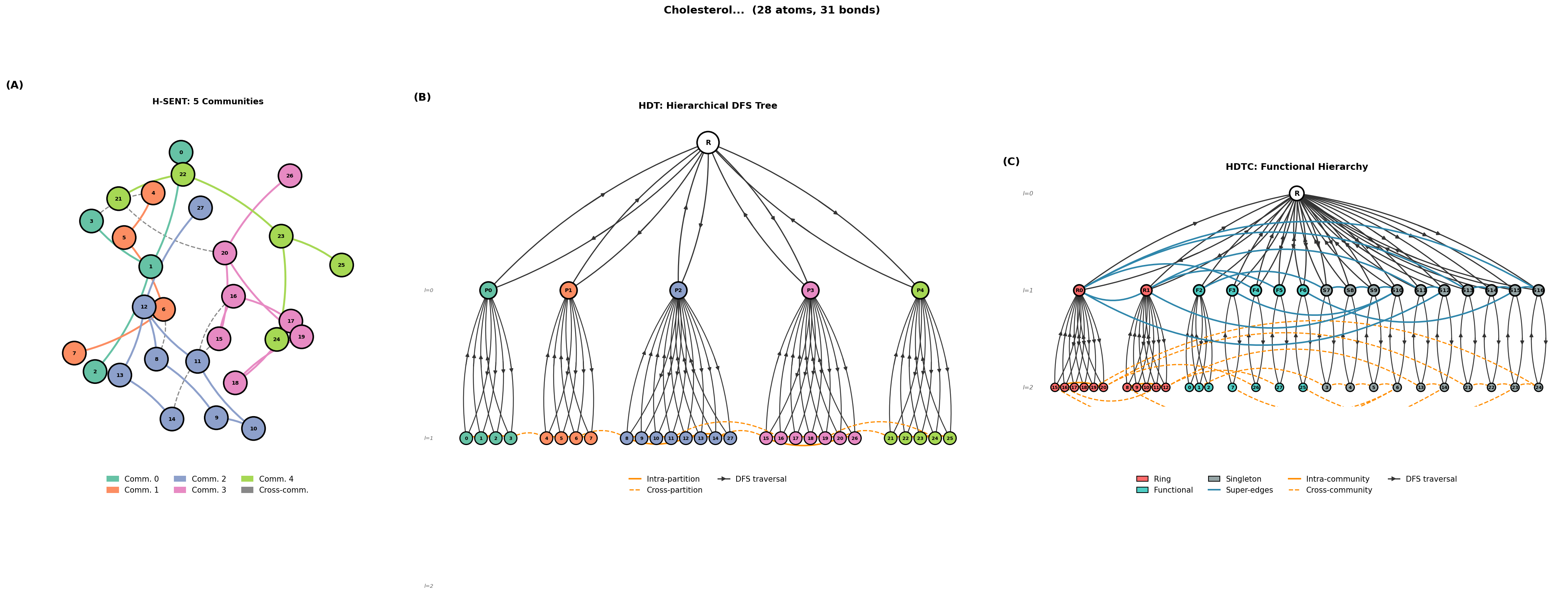
Tokenization Schemes
Given a coarsened hierarchy, we serialize it into a flat token sequence for autoregressive modeling. A shared design principle across all tokenizers is coarse-before-fine ordering: community-level structure always precedes atom-level detail, so the model first commits to the high-level decomposition before filling in local connectivity. H-SENT (Hierarchical SENT) encodes multi-level partition structure with explicit partition boundaries and bipartite cross-community edge blocks; each community’s atoms are serialized via a SENT walk with back-edge brackets. HDT (Hierarchical DFS Tokenization) encodes the full tree structure using depth-first traversal with ENTER/EXIT nesting tokens, capturing both intra- and inter-community edges as back-edges during the DFS, with no separate bipartite reconstruction needed. HDTC (HDT with Composition) is the most expressive scheme: it types each community node (ring, functional group, singleton), encodes a super-graph of inter-community bonds, and serializes atom-level detail within each typed community block.
Results
We evaluate unconditional generation on MOSES (~1M drug-like molecules, 10–26 heavy atoms) and COCONUT (~5K complex natural products, 30–100 heavy atoms, ≥4 rings). All models use a GPT-2 backbone (12 layers, 768 hidden, 12 heads, ~85M parameters) trained with identical hyperparameters. We generate 500 molecules per model and evaluate against 5,000 randomly sampled references from the combined train+test set. We report four representative tokenizers: SENT (flat baseline), H-SENT MC (hierarchical flat-walk), HDT MC (motif-community hierarchy), and HDTC (typed compositional). Bold = best among the four per dataset.
| MOSES (full ref.) | COCONUT (full ref.) | |||||||
|---|---|---|---|---|---|---|---|---|
| SENT | H-SENT MC | HDT MC | HDTC | SENT | H-SENT MC | HDT MC | HDTC | |
| Validitya ↑ | 0.868 | 0.851 | 0.891 | 0.835 | 0.581 | 0.881 | 0.878 | 0.893 |
| Uniquenessa ↑ | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 0.798 | 0.855 | 0.772 |
| Noveltya ↑ | 0.986 | 0.938 | 0.936 | 0.955 | 1.000 | 0.887 | 0.927 | 0.925 |
| FCDb ↓ | 2.35 | 2.44 | 2.30 | 2.44 | 5.40 | 4.60 | 3.25 | 3.72 |
| SNNa ↑ | 0.396 | 0.439 | 0.445 | 0.435 | 0.264 | 0.783 | 0.779 | 0.767 |
| Fraga ↑ | 0.995 | 0.995 | 0.995 | 0.992 | 0.963 | 0.954 | 0.969 | 0.972 |
| Scaffa ↑ | 0.860 | 0.896 | 0.864 | 0.828 | 0.000 | 0.143 | 0.114 | 0.116 |
| IntDiva ↑ | 0.864 | 0.859 | 0.855 | 0.858 | 0.884 | 0.880 | 0.881 | 0.879 |
| PGDc ↓ | 0.000 | 0.000 | 0.000 | 0.051 | 0.308 | 0.317 | 0.000 | 0.125 |
| FG MMDd ↓ | 0.002 | 0.002 | 0.002 | 0.002 | 0.002 | 0.002 | 0.002 | 0.002 |
| SMARTS MMDd ↓ | 0.002 | 0.003 | 0.002 | 0.002 | 0.003 | 0.004 | 0.002 | 0.003 |
| Ring MMDd ↓ | 0.004 | 0.006 | 0.004 | 0.006 | 0.011 | 0.004 | 0.002 | 0.003 |
| BRICSd ↓ | 0.016 | 0.018 | 0.018 | 0.022 | 0.038 | 0.045 | 0.036 | 0.036 |
| Motif Ratee ↑ | 0.790 | 0.832 | 0.806 | 0.836 | 0.575 | 0.647 | 0.645 | 0.708 |
On MOSES (simple drug-like molecules), hierarchy provides modest but consistent improvements: SENT achieves 86.8% validity while HDT MC reaches 89.1% with the best FCD (2.30) and PGD (0.000). H-SENT MC achieves the best scaffold similarity (0.896). HDTC leads on motif rate (0.836). All four achieve near-identical motif-level fidelity (FG MMD ≤ 0.002). On COCONUT (complex natural products), the gap is dramatic: SENT drops to 58.1% validity while all hierarchical tokenizers achieve 87–90%, a 31-point improvement. No single model dominates distributional metrics: H-SENT MC leads SNN (0.783) and scaffold similarity (0.143), HDT MC achieves perfect PGD (0.000) and the best FCD (3.25), and HDTC leads motif rate (0.708) and fragment similarity (0.972). All hierarchical models achieve comparable distributional fidelity (FCD 3.1–4.6, all MMDs ≤ 0.004), suggesting that the choice of tokenizer and coarsening is less about overall quality than about which specific aspects of the distribution to optimize.
Full Results
MOSES Full Results (all 8 model variants)
| SENT | H-SENT MC | H-SENT SC | H-SENT HAC | HDT MC | HDT SC | HDT HAC | HDTC | |
|---|---|---|---|---|---|---|---|---|
| Validity ↑ | 0.868 | 0.851 | 0.842 | 0.209 | 0.891 | 0.220 | 0.213 | 0.835 |
| Uniqueness ↑ | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 |
| Novelty ↑ | 0.986 | 0.938 | 0.811 | 0.876 | 0.936 | 0.882 | 0.859 | 0.955 |
| FCD ↓ | 2.35 | 2.44 | 2.57 | 10.81 | 2.30 | 9.27 | 10.89 | 2.44 |
| SNN ↑ | 0.396 | 0.439 | 0.436 | 0.462 | 0.445 | 0.454 | 0.459 | 0.435 |
| Frag ↑ | 0.995 | 0.995 | 0.993 | 0.951 | 0.995 | 0.962 | 0.950 | 0.992 |
| Scaff ↑ | 0.860 | 0.896 | 0.710 | 0.669 | 0.864 | 0.603 | 0.687 | 0.828 |
| IntDiv ↑ | 0.864 | 0.859 | 0.861 | 0.846 | 0.855 | 0.853 | 0.855 | 0.858 |
| PGD ↓ | 0.000 | 0.000 | 0.045 | 0.710 | 0.000 | 0.719 | 0.763 | 0.051 |
| FG MMD ↓ | 0.002 | 0.002 | 0.002 | 0.006 | 0.002 | 0.005 | 0.006 | 0.002 |
| SMARTS MMD ↓ | 0.002 | 0.003 | 0.003 | 0.013 | 0.002 | 0.007 | 0.013 | 0.002 |
| Ring MMD ↓ | 0.004 | 0.006 | 0.002 | 0.023 | 0.004 | 0.012 | 0.019 | 0.006 |
| BRICS MMD ↓ | 0.016 | 0.018 | 0.020 | 0.061 | 0.018 | 0.055 | 0.061 | 0.022 |
| Motif Rate ↑ | 0.790 | 0.832 | 0.725 | 0.900 | 0.806 | 0.868 | 0.883 | 0.836 |
COCONUT Full Results (all 8 model variants)
| SENT | H-SENT MC | H-SENT SC | H-SENT HAC | HDT MC | HDT SC | HDT HAC | HDTC | |
|---|---|---|---|---|---|---|---|---|
| Validity ↑ | 0.581 | 0.881 | 0.875 | 0.889 | 0.878 | 0.885 | 0.899 | 0.893 |
| Uniqueness ↑ | 1.000 | 0.798 | 0.878 | 0.771 | 0.855 | 0.875 | 0.872 | 0.772 |
| Novelty ↑ | 1.000 | 0.887 | 0.919 | 0.938 | 0.927 | 0.913 | 0.939 | 0.925 |
| FCD ↓ | 5.40 | 4.60 | 3.12 | 4.28 | 3.25 | 3.40 | 3.18 | 3.72 |
| SNN ↑ | 0.264 | 0.783 | 0.771 | 0.747 | 0.779 | 0.777 | 0.782 | 0.767 |
| Frag ↑ | 0.963 | 0.954 | 0.972 | 0.968 | 0.969 | 0.964 | 0.969 | 0.972 |
| Scaff ↑ | 0.000 | 0.143 | 0.141 | 0.125 | 0.114 | 0.151 | 0.115 | 0.116 |
| IntDiv ↑ | 0.884 | 0.880 | 0.882 | 0.881 | 0.881 | 0.880 | 0.880 | 0.879 |
| PGD ↓ | 0.308 | 0.317 | 0.054 | 0.377 | 0.000 | 0.027 | 0.000 | 0.125 |
| FG MMD ↓ | 0.002 | 0.002 | 0.002 | 0.002 | 0.002 | 0.001 | 0.002 | 0.002 |
| SMARTS MMD ↓ | 0.003 | 0.004 | 0.002 | 0.004 | 0.002 | 0.002 | 0.002 | 0.003 |
| Ring MMD ↓ | 0.011 | 0.004 | 0.002 | 0.004 | 0.002 | 0.002 | 0.002 | 0.003 |
| BRICS ↓ | 0.038 | 0.045 | 0.034 | 0.036 | 0.036 | 0.040 | 0.036 | 0.036 |
| Motif Rate ↑ | 0.575 | 0.647 | 0.616 | 0.603 | 0.645 | 0.632 | 0.634 | 0.708 |
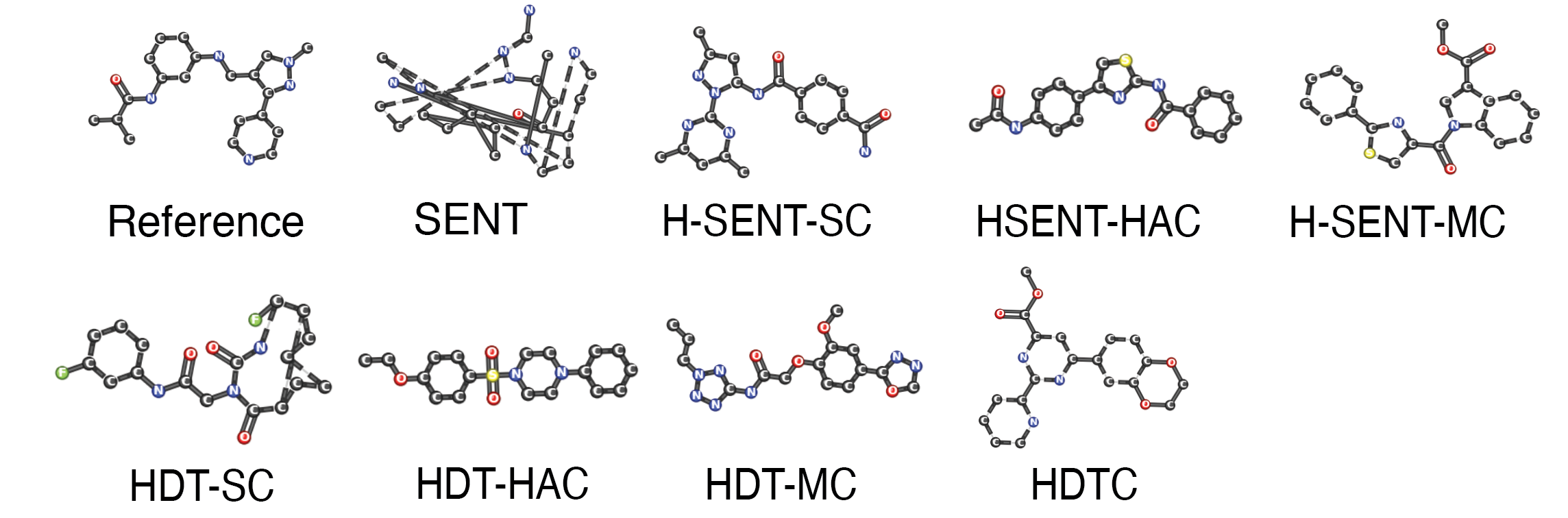
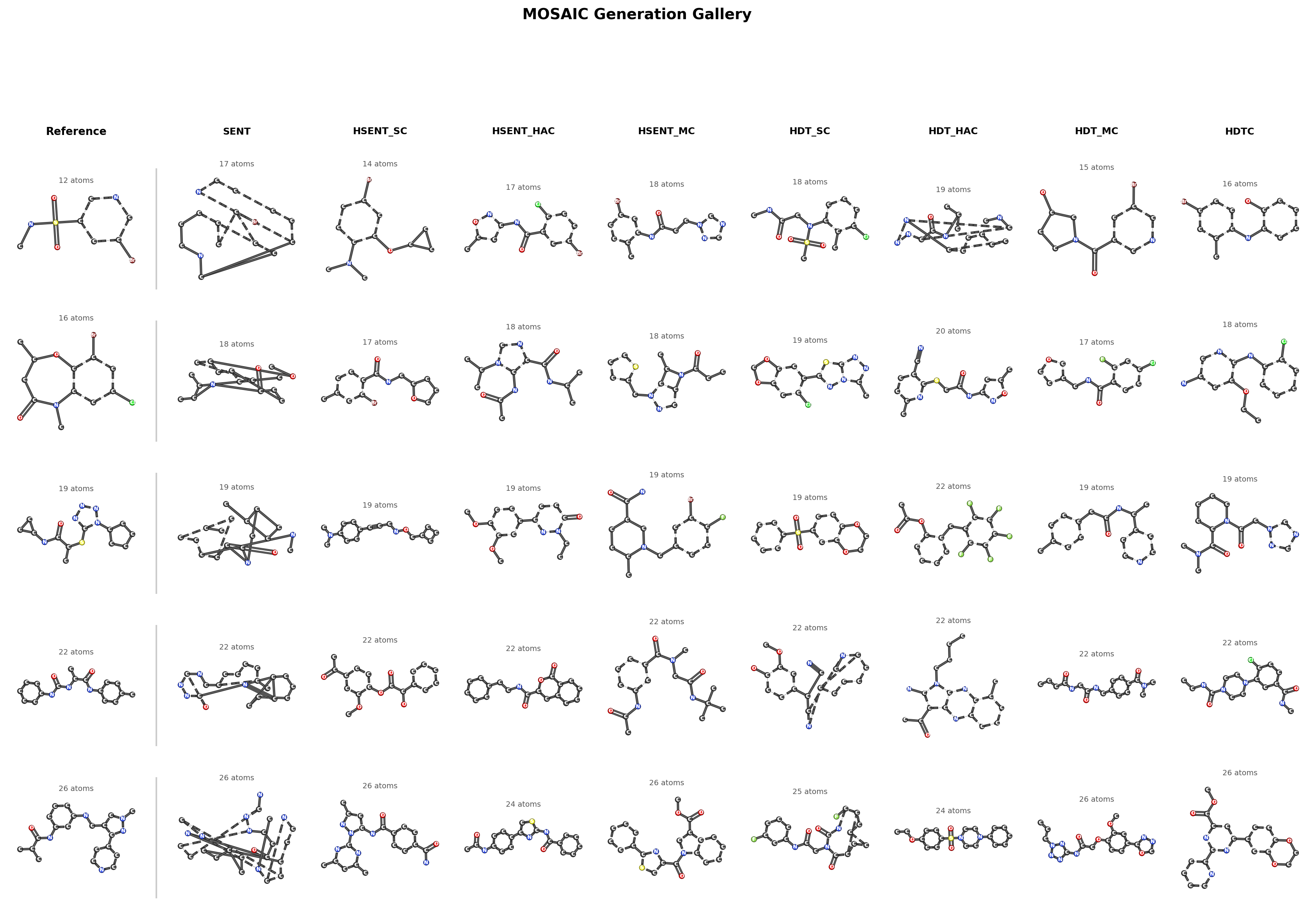
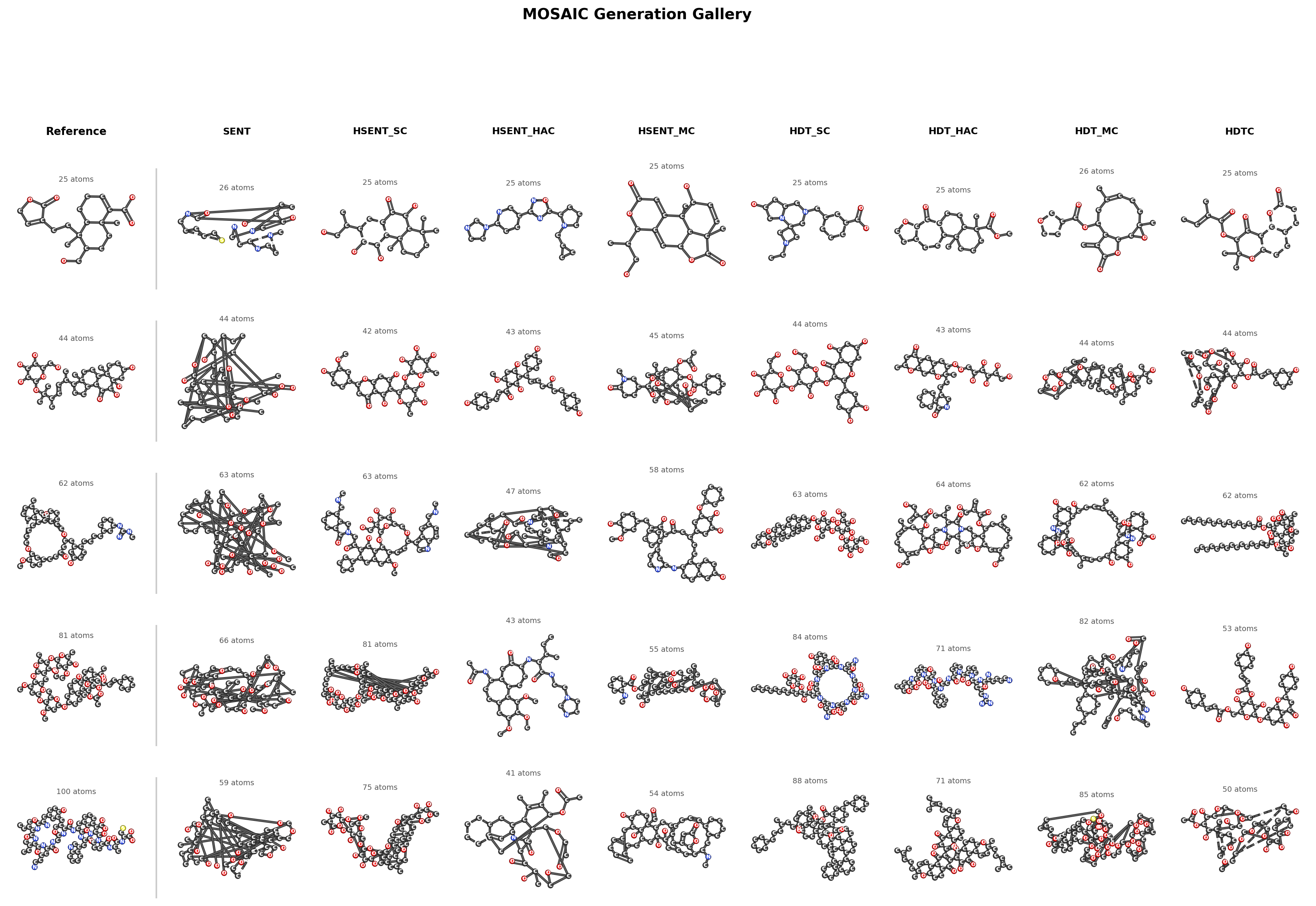
Generation Gallery
Discussion
Hierarchy is necessary for complex molecules. On MOSES (~20 atoms), flat walks achieve 86.8% validity and hierarchy adds only 2.3 points. On COCONUT (30–100 atoms), flat walks drop to 58% validity while all hierarchical tokenizers reach 87–90%, a 31-point gap. The benefit scales with molecular complexity: larger molecules have real compositional structure that hierarchy can exploit.
Enumeration and traversal are complementary. H-SENT serializes molecules as a catalog of parts plus a wiring diagram (explicit index blocks and bipartite edge lists), excelling at distributional precision (best FCD 3.12, best Subst TV/KL). HDT serializes molecules as one continuous DFS walk with nesting, excelling at valid and diverse generation (best validity 89.9%, best PGD 0.000). The coarsening strategy amplifies this: spectral coarsening pairs with H-SENT’s enumeration, HAC pairs with HDT’s traversal.
Coarsening robustness scales with molecular size. On small MOSES molecules, only chemistry-aware MC coarsening works (generic methods drop to ≤22% validity). On complex COCONUT molecules, all coarsening methods achieve 87–90% validity, because larger molecules contain genuine hierarchical motifs that even naive algorithms can discover.
Typed decomposition gives motif precision, not distributional dominance. HDTC’s R/F/S type labels yield the highest motif rate (0.708) and fragment similarity (0.972), but untyped models match or beat it on FCD, SNN, and PGD with greater diversity. The diversity cost of hierarchy is a one-time flat-vs-hierarchical penalty, not a gradient that worsens with more constraint.
Substructure metrics generalize better than molecule-level similarity. SNN and scaffold similarity drop 2–5× between full-reference and test-only evaluation, while fragment similarity and MMD metrics remain stable. This gap is uniform across all models, driven by the small training set (5K molecules) rather than tokenizer-specific memorization.
BibTeX Citation
@article{bian2025mosaic, title = {Beyond Flat Walks: Compositional Abstraction for Autoregressive Molecular Generation}, author = {Bian, Kaiwen and Yang, Andrew H. and Parviz, Ali and Mishne, Gal and Wang, Yusu}, year = {2025}, url = {https://github.com/KevinBian107/MOSAIC},}